Genes2Networks is a command-line software tool that can be used to place lists of mammalian genes in the context of a background mammalian signalome and interactome networks.
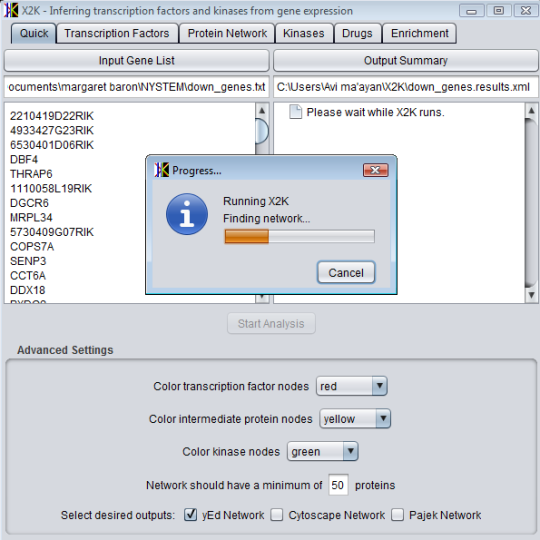
The input to the program is a list of human Entrez Gene gene symbols and background networks in SIG format, while the output includes: all identified interactions for the genes/proteins, a subnetwork connecting the genes/proteins using intermediate components that are used to connect the genes, ranking of the specificity of intermediate components to interact with the list of genes/proteins, and a clustering analysis of the genes/proteins from the seed list based on their distance from one another in network space.
Requirements:
DOS, Windows 3.x/95/98/Me/NT/2000/XP/2003 Server/Vista

Comments not found